Multivariate morphometrics provide a suite of tools to describe complex biological shapes, and have been essential to recent developments in analyses of trait diversification. However, most multivariate morphometric methods (both linear and geometric morphometric) are intolerant of missing data. Missing data from broken, incomplete, distorted or otherwise damaged specimens can be dealt with in one of three ways (or a combination of):
- Remove that measurement from all specimens
- Remove the incomplete specimen from the dataset
- Estimate the missing data (or accommodate missing data with some other statistical method)

Option 1 may be acceptable if all missing data is restrict to one (or a few) traits (linear measurements or geometric morphometric landmarks) that are unlikely to have a strong impact on overall shape, but more practically often limits datasets to just a small number of traits. Option 2 may be acceptable if few specimens are damaged, and are from species/populations/etc. represented by many other individuals in the dataset. However, incomplete specimens can often represent fossil lineages or rare taxa that are poorly represented in collections, and in this case even damaged specimens may need to be utilized to fully capture the morphological variation within a clade. Fossils in particular are critical in understanding the evolution of biological shape.
As part of collaborations with Dr. Caleb Brown and Dr. Don Jackson, I helped to evaluate the effectiveness of several missing data estimation techniques within linear morphometric analyses (Brown et al. 2012). Additionally, I developed several approaches for incorporating bias into the distribution of missing morphometric data. While many studies select data at random to simulate missing data, in reality missing data may be biased in several ways. For example, missing measurements are likely to be anatomically clustered if damage is constrained to one area (see fossil image to the left). Also, rare taxa (with few specimens to select from) may be more likely to be incomplete, compared to common species where a large number of complete individuals may be selected from. We found that the impact of these biases varied across a linear morphometric dataset taken from crocodilian crania.
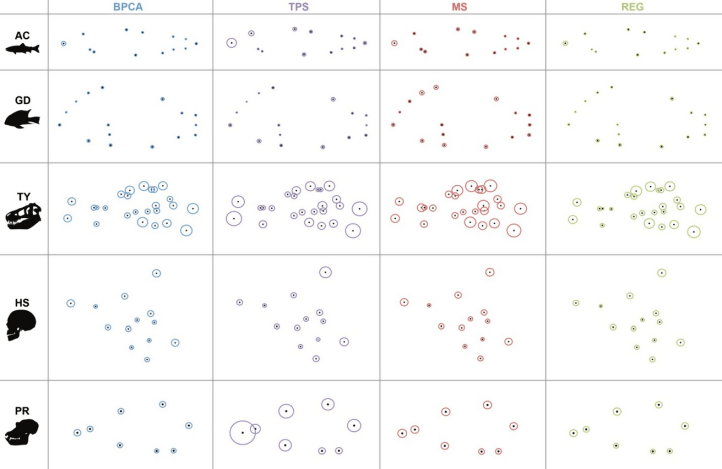
In a follow up study, I compared the impact of missing data estimation to the exclusion of incomplete specimens (option 2 above) in the variation of shape data in geometric morphometrics. Across several datasets varying in size and taxonomic composition, I found that the inclusion of incomplete specimens was almost always preferred over their removal in the analysis of shape, but that the effectiveness of different estimation methods varied both across and within datasets. Interestingly, one of the most common approaches in missing data estimation within geometric morphometrics, thin plate spline interpolation (TPS) was one of the least reliable across datasets. In this paper I made several recommendations on how to use the dataset of complete specimens to evaluate different methods using simulation approaches prior to missing data estimation.
All statistical functions for these two studies were made available through the R package LOST. Recently I have been working on updating LOST to better accommodate 3D data, and to make it easier to move data between LOST and common R packages for geometric morphometrics like “geomorph”. Stay tuned for updates!
Selected Publications
Arbour, J.H. and Brown, C.M.. 2014. Incomplete specimens in geometric morphometric analyses. Methods in Ecology and Evolution 5(1):16-26. (link)
Brown, C. M., Arbour, J.H. & Jackson, D. 2012. Testing of the effect of missing data estimation and distribution in morphometric multivariate data analyses. Systematic Biology 61: 941-954. (link)